We are very proud that this July release sees the introduction of some exceptional new functionality on the VarSome platform. These changes reflect our company mission statement “To enable anyone to find, share, use the most comprehensive human genome data - and to collaborate to improve health and lives around the world.”
Thank you as always for the continued feedback and support - and welcome to VarSome 10.0
VarSome 10.0
Automated ACMG classification for CNVs
In VarSome Clinical the major update in this release is the introduction of CNV classifications. In addition to the ability to call CNVs directly from FASTQ, you can now launch CNV VCFs and automatically classify variants using the ACMG CNV classification guidelines.
For further details please see the release notes for the CNV ACMG Classifier.
CNV Databases
VarSome’s structural variant browser has been improved and incorporates a number of new databases:
The following databases are now visible in both VarSome (and VarSome Premium) and VarSome Clinical:
- Decipher
- Clinvar CNVs
- Exac
- DGV
- dbVar
- ClinGen Structural Variants
- gnomAD Structural Variants
Publications
We are proud to announce two major changes to the publication browser in VarSome:
- VarSome AI now automatically identifies variants mentioned in the title & abstract of publications. This will be extended to full text search in future.
- Users can now ‘up vote’ links & classifications provided by other users, as well as comment on them.
For further details please see the release notes for the Publications Browser.
ACMG & AMP
Improvement to rule BP4
Because many of the in-silico predictors are not very good around splice-sites, rule BP4 is now disabled if a variant is both PVS1 and predicted splicing using scSNV. This results in a very small reduction in false negatives with no increase in false positives.
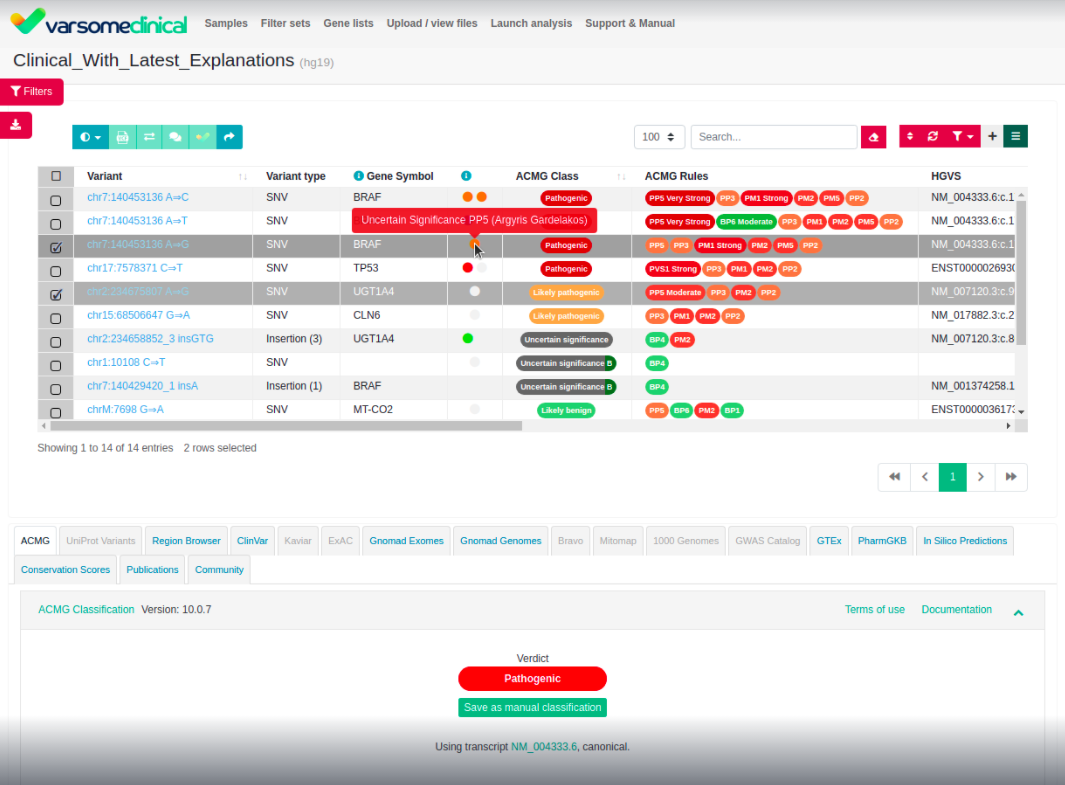
Uncertain Significance
We have enhanced the display of Variants of Unknown Significance (VUS) in VarSome and Clinical to make it clear whether the variant has supporting pathogenic or benign evidence. This is purely a display feature and has no impact on the classification itself, however it may help clinicians better identify edge-cases or variants with conflicting evidence.
This feature is also visible in VarSome Clinical:
GnomAD Mitochondrial variants
We have now incorporated the GnomAD 3.1 Mitochondrial database into our ACMG classifier - this helps improve specificity. Note that there is at present no visualiser for mitochondrial variant frequencies in VarSome.
Premium Databases
Free users will be alerted to the existence of additional Premium data in VarSome which may impact the classification of the variants, this may include data from Cosmic, JAX-CKB, CADD, Polyphen etc.
VarSome Clinical
AMP JAX CKB
JAX CKB is now mandatory for AMP classification: if JAX has been disabled by your site administrator, the AMP classifier will not be used for somatic samples and the corresponding columns will not be visible in reports.
Reporting of Custom Classifications
If you add a custom classification to a variant, it will now be displayed in the report instead of the default automated ACMG classification:
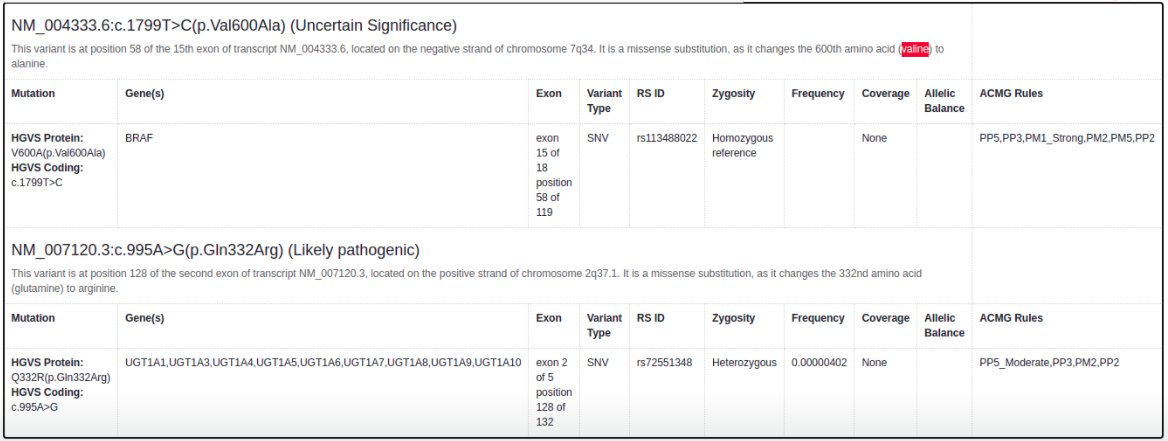
Allelic Balance
The allelic balance (VAF) value has been added to clinical reports:
Publications
The formatting of cited publications has been improved:
Analysis Summary
This report has been improved and extraneous information removed:
VarSome Improvements
Clinical Trials: AACT (Premium)
We have made the AACT component more useful for clinicians. Additional fields have been added that include dates and location and how old the study is. We have also added additional filtering such as by recruitment phase and given drugs.
Community Connection Requests
We have added an additional two columns to our community page that show the number of connection requests and the number of connections already made. It is possible to sort by either of these fields.
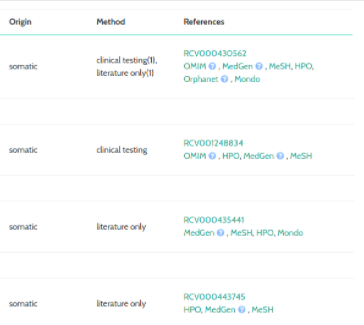
ClinVar & MONDO
The ClinVar component now links directly to the MONDO diseases database:
Further Information and Support
Not already a VarSome Premium or VarSome Clinical user? Get in touch and ask for a free trial.
As ever we hope you find these changes and improvements helpful, we’d love to hear any suggestions you may have, support is available as usual from support@varsome.com
- The VarSome Team


Submit a Comment