We are very proud to launch our CNV ACMG Classification as part of VarSome 10.0. This note details the new CNV functionality we have introduced as part of VarSome Clinical.
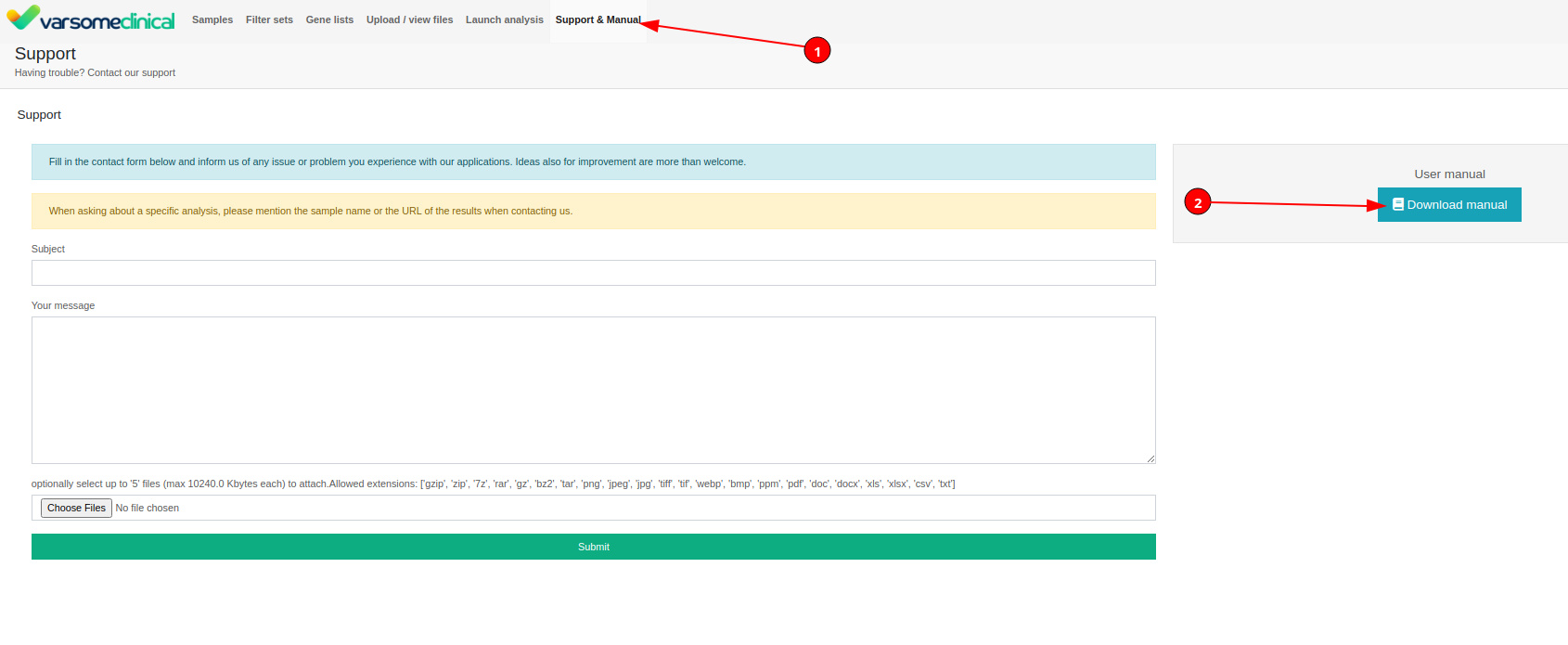
For further detail on how to use all CNV functionality, VarSome Clinical users should refer to our training manual. This can be accessed through the VarSome Clinical portal by selecting 1) Support & Manual, and 2) Download Manual.
CNV annotation from VCF
VarSome Clinical now includes a pipeline to annotate CNVs/SV from VCFs. You can provide a VCF containing CNV/SV variants when launching a new analysis either from FASTQ or VCF. Additionally, you can run a CNV sub-analysis from an already analyzed sample.
CNV ACMG Classification
The columns containing the ACMG classification of CNVs and the set of triggered ACMG rules are now displayed in the CNV variant table of VarSome Clinical. Results are sorted by decreasing pathogenicity.
CNV Structural Variant browser
Several new CNV related databases have now been added to our structural variant browser for CNV analysis in VarSome. These include Decipher, Clinvar CNVs, Exac, DGV, DBVar, ClinGen Regions, ClinGen CNVs and gnomAD. You can see the complete list of data sources available here.
In VarSome Clinical, we filter the CNVs reported at the CNV region to retain the CNVs that are relevant for the classification. The filtered CNVs can be accessed from the “Known CNVs” tab below the CNV variant table.
Sensitive Mode for CNV calling
A “Sensitive mode” for non-WGS CNV analyses is now available in VarSome Clinical. This mode uses a lower threshold for detecting CNVs compared to the CNV standard mode, resulting in more sensitive calling and typically in a higher number of calls. You can enable this feature for either somatic or germline samples (or both) in the group settings. Please note that these settings are only available to the group administrator and any changes will be applied to all users of the group.
CNV Filtering
We have added a number of new filtering options for CNV results. You can now filter by the total number of genes or exons overlapping the CNV region. You can also filter by CNV frequency (calculated from gnomAD database), reads expected, observed, and the ratio between both of them.
CNVs Export
CNVs data is now included in the export of data to excel.
Further Information and Support
Not already a VarSome Clinical user? Get in touch and ask for a free trial.
As ever we hope you find these changes and improvements helpful, we’d love to hear any suggestions you may have, support is available as usual from support@varsome.com
- The VarSome Team


Submit a Comment